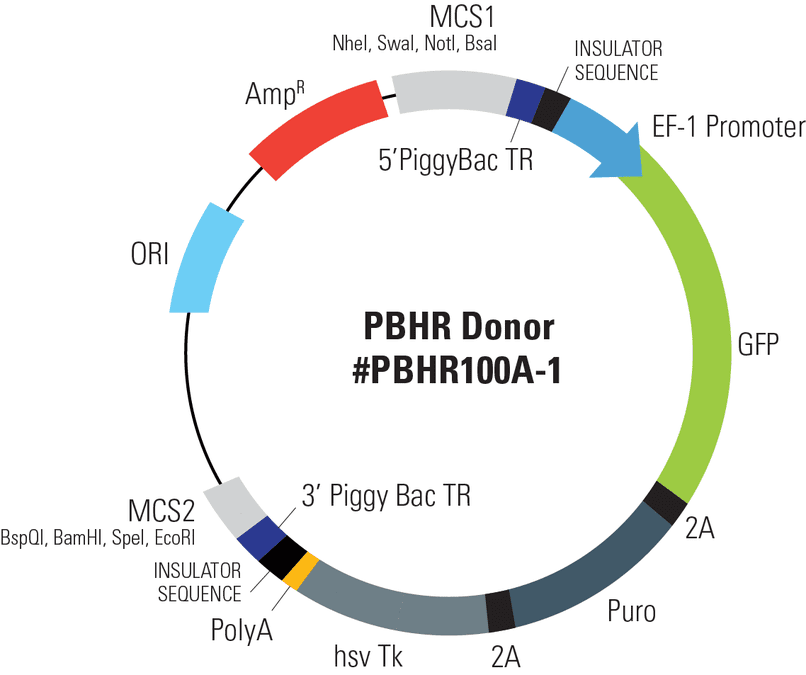
PiggyBac Gene Editing HR Targeting Vector (MCS1-5’PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3′ PB TR-MCS2)
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| PBHR100A-1 | piggyBac-HR with GFP+Puro markers and TK selection (MCS1-5'PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3′ PB TR-MCS2) for Gene Editing | 10 µg | $1660 |
|
||||
Overview
Overview
Seamless gene editing
Use the PiggyBac Gene Editing HR Targeting Vector (MCS1-5’PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3′ PB TR-MCS2) to get seamless gene editing—leave no trace of vector sequences behind.
This special HR Donor works with the Excision-only PiggyBac Transposase instead of the Cre-LoxP system. Simply proceed with your CRISPR/Cas9 gene editing as usual—clone your homology arms into MCS1 and MCS2, use dual GFP and puromycin selection to find integrants, and enrich for on-target events using negative thymidine kinase (TK) selection (Figure 1).
Why use an HR targeting vector?
Even though gene knock-outs can result from DSBs caused by Cas9 alone, SBI recommends the use of HR targeting vectors (also called HR donor vectors) for more efficient and precise mutation. HR donors can supply elements for positive or negative selection ensuring easier identification of successful mutation events. In addition, HR donors can include up to 6-8 kb of open reading frame for gene knock-ins or tagging, and, when small mutations are included in either 5’ or 3’ homology arms, can make specific, targeted gene edits.
Choose the right HR Targeting Vector for your project
| Catalog # | HR Donor Vector | Features* | Application |
|||
|---|---|---|---|---|---|---|
| Gene | Gene | Gene Edits | Gene Tagging | |||
| HR100PA-1 | MCS1-LoxP-MCS2-MCS3-pA-LoxP-MCS4 | Basic HR Donor | ||||
| HR110PA-1 | MCS1-EF1α-RFP-T2A-Puro-pA-MCS2 | Removable RFP marker and puromycin selection | ||||
| HR120PA-1 | GFP-pA-LoxP-EF1α-RFP-T2A-Puro-pA-LoxP-MCSPuro-pA-LoxP-MCS | Tag with GFP fusion Removable RFP marker and puromycin selection | ||||
| HR130PA-1 | T2A-GFP-pA-loxP-EF1α-RFP-T2A-Puro-pA-LoxP-MCSA-loxP-EF1α-RFP-T2A-Puro-pA-LoxP-MCS | Co-express GFP with “tagged” gene via T2A Removable RFP marker and puromycin selection | ||||
| HR150PA-1 | GFP-T2A-Luc-pA-loxP-EF1α-RFP-T2A-Puro-pA-LoxP-MCS | Tag with GFP fusion and co-express luciferase via T2A Removable RFP marker and puromycin selection | ||||
| HR180PA-1 | IRES-GFP-pA-loxP-MCS1-EF1α-RFP-T2A-Puro-pA-LoxP-MCS2 | Co-express GFP with “tagged” gene via IRES Removable RFP marker and puromycin selection | ||||
| HR210PA-1 | MCS1-LoxP-EF1α-GFP-T2A-Puro-P2A-hsvTK-pA-LoxP-MCS2 | Removable GFP marker, puromycin selection, and TK selection | ||||
| HR220PA-1 | GFP-pA-LoxP-EF1α-RFP-T2A-Hygro-pA-LoxP-MCS | Tag with GFP fusion Removable RFP ,arker and hygromycin Selection | ||||
| HR410PA-1 | MCS1-EF1α-GFP-T2A-Puro-pA-MCS2 | Removable GFP marker and puromycin selection | ||||
| HR510PA-1 | MCS1-EF1α-RFP-T2A-Hygro-pA-MCS2 | Removable RFP marker and hygromycin selection | ||||
| HR700PA-1 | MCS1-EF1α-GFP-T2A-Puro-pA-MCS2-PGK-hsvTK | Enrich for on-target integration with negative TK selection** Removable GFP marker and puromycin selection | ||||
| HR710PA-1 | MCS1-EF1α-RFP-T2A-Hygro-pA-MCS2-PGK-hsvTK | Enrich for on-target integration with negative TK selection** Removable RFP marker and hygromycin selection | ||||
| HR720PA-1 | MCS1-EF1α-Blasticidin-pA-MCS2-PGK-hsvTK | Enrich for on-target integration with negative TK selection** Removable blasticidin selection | ||||
| GE602A-1 | pAAVS1D-PGK-MCS-EF1α-copGFPpuro | First generation AAVS1-targeting HR Donor | ||||
| GE603A-1 | pAAVS1D-CMV-RFP-EF1α-copGFPpuro | First generation AAVS1-targeting HR Donor (positive control for GE602A-1) | ||||
| GE620A-1 | AAVS1-SA-puro-MCS | Second generation AAVS1-targeting HR Donor Promoterless to knock-in any gene or promoter-gene combination | ||||
| GE622A-1 | AAVS1-SA-puro-EF1α-MCS | Second generation AAVS1-targeting HR Donor Constitutive expression of your gene-of-interest | ||||
| GE624A-1 | AAVS1-SA-puro-MCS-GFP | Second generation AAVS1-targeting HR Donor Create reporter cell lines | ||||
| CAS620A-1 | AAVS1-SA-puro-EF1α-hspCas9 | Knock-in Cas9 to the AAVS1 site | ||||
| MCS1-5'PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3' PB TR-MCS2 | Use with the PiggyBac Transposon System Enables seamless gene editing with no residual footprint (i.e. completely remove vector sequences) | |||||
| *All HR Target Vectors except PBHR100A-1 contain LoxP sites. Any sequences that are integrated between the two LoxP sites can be removed through transient expression of Cre Recombinase. **The clever design of these HR Donors enables enrichment for on-target integration events. A PGK-hsvTK cassette is included outside of the homology arms. Because of this configuration, on-target integration that results from homologous recombination will not include the PGK-hsvTK cassette—only randomly-integrated off-target events will lead to integration of PGK-hsvTK and resulting TK activity. Therefore, TK selection will negatively select against off-target integrants. Click on any one of these vectors to see a diagram of how the negative selection works. |
||||||
* SBI is fully licensed to distribute PiggyBac vectors as a partnership with Hera BioLabs, Inc.
References
How It Works
How It Works
Getting seamless gene editing with the Excision-only PiggyBac Transposase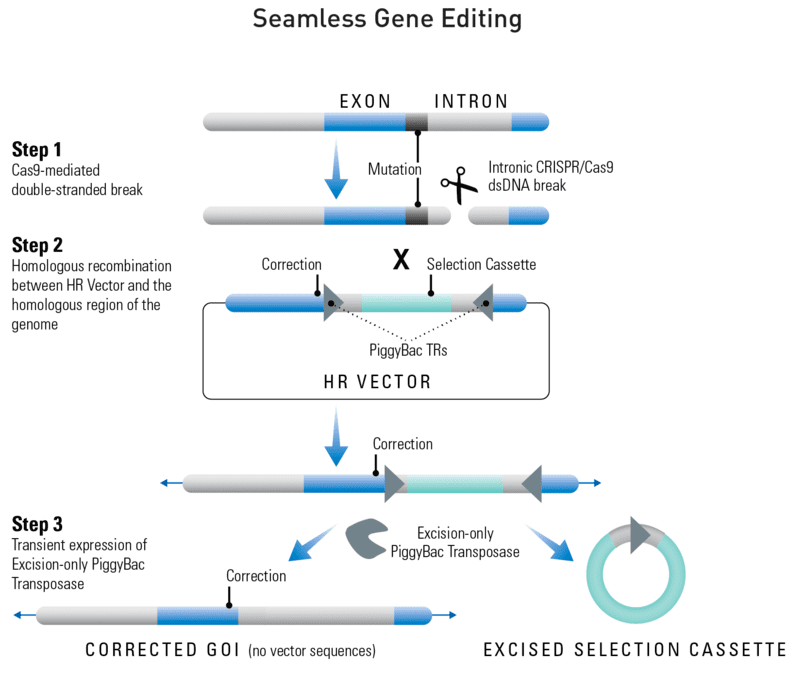
Figure 1. Seamless gene editing with the Excision-only PiggyBac Transposase. Step 1: Cas9 creates a double-stranded break (DSB) in the genomic DNA at a site that is complimentary to the gRNA. For gene editing, this DSB should be within an intron. Step 2: The DNA repair machinery is recruited to the DSB. In the presence of an HR Donor with homology to the region adjacent to the DSB (blue areas of the genomic and vector DNA) homologous recombination (HR) is favored over non-homologous end joining (NHEJ). If one of the homology arms of the HR donor contains the gene edit, it will be incorporated into the gene through the HR repair process. Step 3: Like the Cre-LoxP system, the PiggyBac Transposon system relies on an enzyme—the transposase—to mediate a site-specific recombination event between two sites, the 5’ PiggyBac Terminal Repeat (TR) and the 3’ PiggyBac TR. However, unlike the Cre-LoxP system, the Excision-only PiggyBac Transposase completely removes the 5’ and 3’ PiggyBac TRs as well as any sequences in between the two TRs. The design of the PiggyBac Gene Editing HR Targeting Vector (MCS1-5’PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3′ PB TR-MCS2) places two homology arms just outside of the PiggyBac Terminal Repeats (TRs). Thus, after Excision-only PiggyBac Transposase activity, all vector sequences are removed.
Genome engineering with CRISPR/Cas9For general guidance on using CRISPR/Cas9 technology for genome engineering, including the design of HR Targeting Vectors, take a look at our CRISPR/Cas9 tutorials as well as the following application notes:
CRISPR/Cas9 Gene Knock-Out Application Note (PDF) »CRISPR/Cas9 Gene Editing Application Note (PDF) »
CRISPR/Cas9 Gene Tagging Application Note (PDF) »
Supporting Data
FAQs
Documentation
Citations
Related Products
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| PBHR100A-1 | piggyBac-HR with GFP+Puro markers and TK selection (MCS1-5'PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3′ PB TR-MCS2) for Gene Editing | 10 µg | $1660 |
|
||||
Overview
Overview
Seamless gene editing
Use the PiggyBac Gene Editing HR Targeting Vector (MCS1-5’PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3′ PB TR-MCS2) to get seamless gene editing—leave no trace of vector sequences behind.
This special HR Donor works with the Excision-only PiggyBac Transposase instead of the Cre-LoxP system. Simply proceed with your CRISPR/Cas9 gene editing as usual—clone your homology arms into MCS1 and MCS2, use dual GFP and puromycin selection to find integrants, and enrich for on-target events using negative thymidine kinase (TK) selection (Figure 1).
Why use an HR targeting vector?
Even though gene knock-outs can result from DSBs caused by Cas9 alone, SBI recommends the use of HR targeting vectors (also called HR donor vectors) for more efficient and precise mutation. HR donors can supply elements for positive or negative selection ensuring easier identification of successful mutation events. In addition, HR donors can include up to 6-8 kb of open reading frame for gene knock-ins or tagging, and, when small mutations are included in either 5’ or 3’ homology arms, can make specific, targeted gene edits.
Choose the right HR Targeting Vector for your project
| Catalog # | HR Donor Vector | Features* | Application |
|||
|---|---|---|---|---|---|---|
| Gene | Gene | Gene Edits | Gene Tagging | |||
| HR100PA-1 | MCS1-LoxP-MCS2-MCS3-pA-LoxP-MCS4 | Basic HR Donor | ||||
| HR110PA-1 | MCS1-EF1α-RFP-T2A-Puro-pA-MCS2 | Removable RFP marker and puromycin selection | ||||
| HR120PA-1 | GFP-pA-LoxP-EF1α-RFP-T2A-Puro-pA-LoxP-MCSPuro-pA-LoxP-MCS | Tag with GFP fusion Removable RFP marker and puromycin selection | ||||
| HR130PA-1 | T2A-GFP-pA-loxP-EF1α-RFP-T2A-Puro-pA-LoxP-MCSA-loxP-EF1α-RFP-T2A-Puro-pA-LoxP-MCS | Co-express GFP with “tagged” gene via T2A Removable RFP marker and puromycin selection | ||||
| HR150PA-1 | GFP-T2A-Luc-pA-loxP-EF1α-RFP-T2A-Puro-pA-LoxP-MCS | Tag with GFP fusion and co-express luciferase via T2A Removable RFP marker and puromycin selection | ||||
| HR180PA-1 | IRES-GFP-pA-loxP-MCS1-EF1α-RFP-T2A-Puro-pA-LoxP-MCS2 | Co-express GFP with “tagged” gene via IRES Removable RFP marker and puromycin selection | ||||
| HR210PA-1 | MCS1-LoxP-EF1α-GFP-T2A-Puro-P2A-hsvTK-pA-LoxP-MCS2 | Removable GFP marker, puromycin selection, and TK selection | ||||
| HR220PA-1 | GFP-pA-LoxP-EF1α-RFP-T2A-Hygro-pA-LoxP-MCS | Tag with GFP fusion Removable RFP ,arker and hygromycin Selection | ||||
| HR410PA-1 | MCS1-EF1α-GFP-T2A-Puro-pA-MCS2 | Removable GFP marker and puromycin selection | ||||
| HR510PA-1 | MCS1-EF1α-RFP-T2A-Hygro-pA-MCS2 | Removable RFP marker and hygromycin selection | ||||
| HR700PA-1 | MCS1-EF1α-GFP-T2A-Puro-pA-MCS2-PGK-hsvTK | Enrich for on-target integration with negative TK selection** Removable GFP marker and puromycin selection | ||||
| HR710PA-1 | MCS1-EF1α-RFP-T2A-Hygro-pA-MCS2-PGK-hsvTK | Enrich for on-target integration with negative TK selection** Removable RFP marker and hygromycin selection | ||||
| HR720PA-1 | MCS1-EF1α-Blasticidin-pA-MCS2-PGK-hsvTK | Enrich for on-target integration with negative TK selection** Removable blasticidin selection | ||||
| GE602A-1 | pAAVS1D-PGK-MCS-EF1α-copGFPpuro | First generation AAVS1-targeting HR Donor | ||||
| GE603A-1 | pAAVS1D-CMV-RFP-EF1α-copGFPpuro | First generation AAVS1-targeting HR Donor (positive control for GE602A-1) | ||||
| GE620A-1 | AAVS1-SA-puro-MCS | Second generation AAVS1-targeting HR Donor Promoterless to knock-in any gene or promoter-gene combination | ||||
| GE622A-1 | AAVS1-SA-puro-EF1α-MCS | Second generation AAVS1-targeting HR Donor Constitutive expression of your gene-of-interest | ||||
| GE624A-1 | AAVS1-SA-puro-MCS-GFP | Second generation AAVS1-targeting HR Donor Create reporter cell lines | ||||
| CAS620A-1 | AAVS1-SA-puro-EF1α-hspCas9 | Knock-in Cas9 to the AAVS1 site | ||||
| MCS1-5'PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3' PB TR-MCS2 | Use with the PiggyBac Transposon System Enables seamless gene editing with no residual footprint (i.e. completely remove vector sequences) | |||||
| *All HR Target Vectors except PBHR100A-1 contain LoxP sites. Any sequences that are integrated between the two LoxP sites can be removed through transient expression of Cre Recombinase. **The clever design of these HR Donors enables enrichment for on-target integration events. A PGK-hsvTK cassette is included outside of the homology arms. Because of this configuration, on-target integration that results from homologous recombination will not include the PGK-hsvTK cassette—only randomly-integrated off-target events will lead to integration of PGK-hsvTK and resulting TK activity. Therefore, TK selection will negatively select against off-target integrants. Click on any one of these vectors to see a diagram of how the negative selection works. |
||||||
* SBI is fully licensed to distribute PiggyBac vectors as a partnership with Hera BioLabs, Inc.
References
How It Works
How It Works
Getting seamless gene editing with the Excision-only PiggyBac Transposase
Figure 1. Seamless gene editing with the Excision-only PiggyBac Transposase. Step 1: Cas9 creates a double-stranded break (DSB) in the genomic DNA at a site that is complimentary to the gRNA. For gene editing, this DSB should be within an intron. Step 2: The DNA repair machinery is recruited to the DSB. In the presence of an HR Donor with homology to the region adjacent to the DSB (blue areas of the genomic and vector DNA) homologous recombination (HR) is favored over non-homologous end joining (NHEJ). If one of the homology arms of the HR donor contains the gene edit, it will be incorporated into the gene through the HR repair process. Step 3: Like the Cre-LoxP system, the PiggyBac Transposon system relies on an enzyme—the transposase—to mediate a site-specific recombination event between two sites, the 5’ PiggyBac Terminal Repeat (TR) and the 3’ PiggyBac TR. However, unlike the Cre-LoxP system, the Excision-only PiggyBac Transposase completely removes the 5’ and 3’ PiggyBac TRs as well as any sequences in between the two TRs. The design of the PiggyBac Gene Editing HR Targeting Vector (MCS1-5’PB TR-EF1α-GFP-T2A-Puro-T2A-hsvTK-pA-3′ PB TR-MCS2) places two homology arms just outside of the PiggyBac Terminal Repeats (TRs). Thus, after Excision-only PiggyBac Transposase activity, all vector sequences are removed.
Genome engineering with CRISPR/Cas9For general guidance on using CRISPR/Cas9 technology for genome engineering, including the design of HR Targeting Vectors, take a look at our CRISPR/Cas9 tutorials as well as the following application notes:
CRISPR/Cas9 Gene Knock-Out Application Note (PDF) »CRISPR/Cas9 Gene Editing Application Note (PDF) »
CRISPR/Cas9 Gene Tagging Application Note (PDF) »