pGreenFire 2.0 - signalling pathway reporters re-engineered for reliable stable cell line generation
NEW PRODUCT ALERT!
SBI is excited to announce the launch of our next generation of pGreenFire signalling pathway reporters! We've upgraded these popular lentivectors with a clever design that enables reliable generation of stable cell lines and have also replaced the conventional luciferase reporter with red firefly luciferase, which opens up the possibility of performing a dual-spectral luciferase assay and delivers greater sensitivity for in vivo applications than conventional luciferase.
pGreenFire Basics
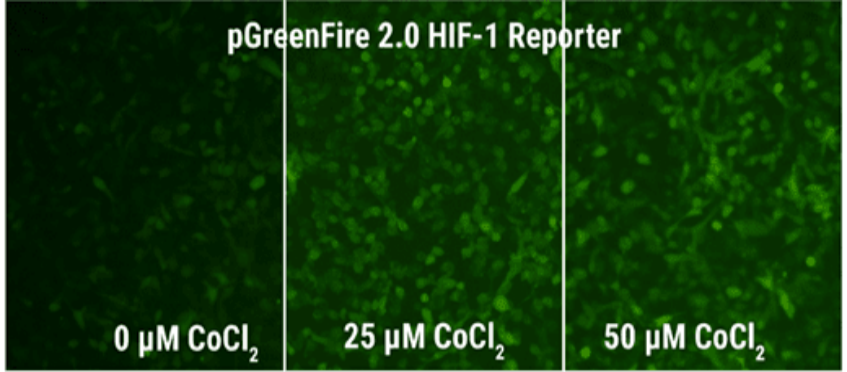
If you're not familiar with our pGreenFire reporters, both 1.0 and 2.0 share a similar core functionality—transcriptional response elements (TREs) are placed upstream of a minimal CMV promoter (mCMV) and the pGreenFire luciferase-T2A-GFP co-expression cassette. In the absence of transcriptional activation, the mCMV promoter has negligible activity resulting in little- to no- luciferase activity or GFP fluorescence (Figure 1). However, upon activation of the TREs, such as in response to the addition of an inducer, the TREs plus the mCMV promoter drive expression of both luciferase and GFP in a dose-dependent fashion (Figure 1). The result is the ability to quantitatively measure pathway activation using luciferase activity or while imaging using GFP.
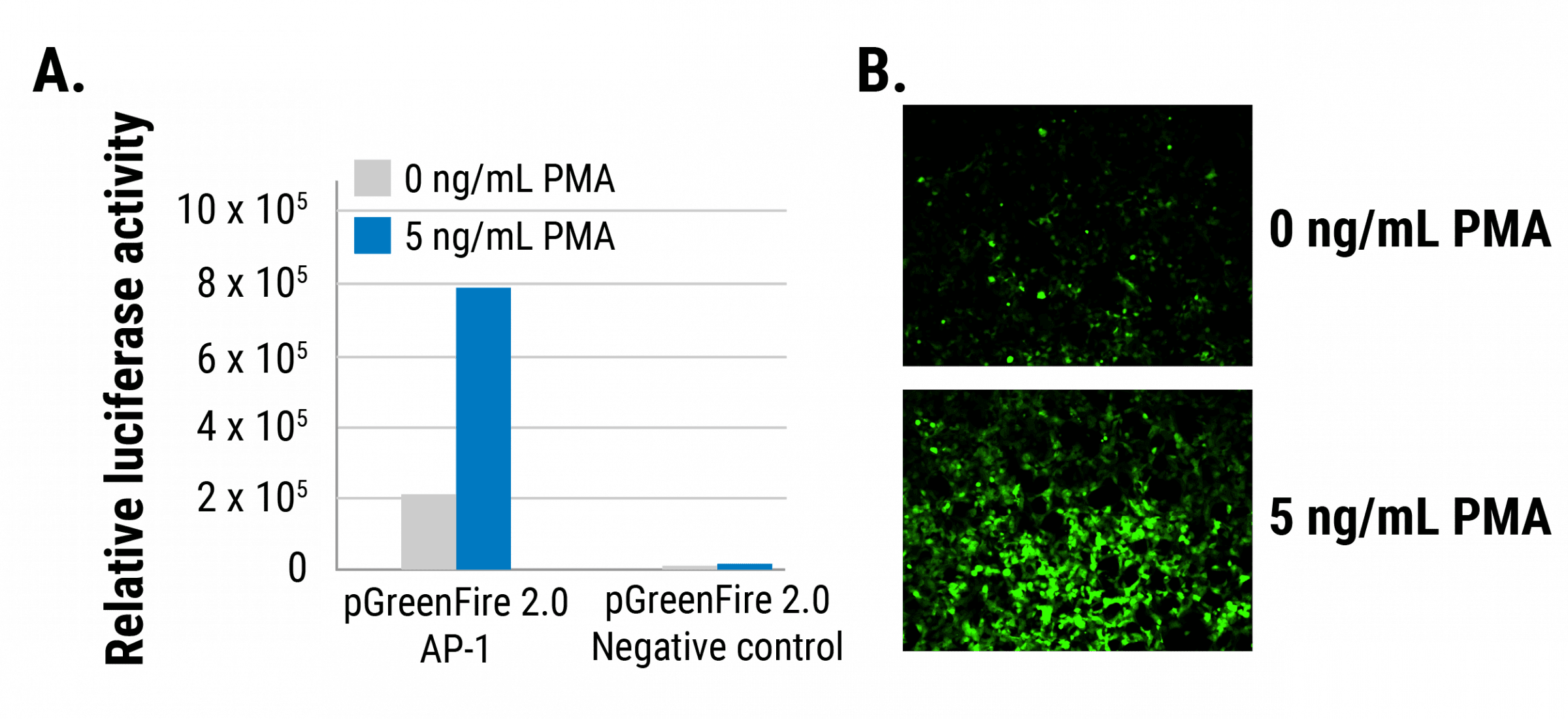
Figure 1. The pGreenFire 2.0 AP-1 Reporter efficiently and quantitatively reports on AP-1 activity in 293FT cells. Relative luciferase activity (A) and GFP activity (B) both increase in response to PMA, an AP-1inducer. Induction is specific to the AP-1 transcription response elements as the pGreenFire 2.0 mCMV Negative Control displays very little luciferase activity regardless of the presence of inducer.
Why pGreenFire 2.0 Lentivectors are better than other signalling pathway reporters
What makes our next-gen pGreenFire 2.0 vectors even better than other TRE reporter vectors is the smart design, which adds in a constitutive selection cassette for stable cell line generation while minimizing interference from the upstream TRE. By using a weak/moderate mPGK promoter to drive the antibiotic selection marker (puromycin resistance) and carefully arranging the conditional reporter genes, the selection marker is reliably expressed without compromising conditional expression of rFLuc and GFP. This means that every pGreenFire 2.0 Lentivector can be used to create stable reporter cell lines.
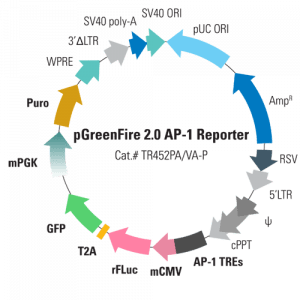
Available pGreenFire 2.0 Lentivectors
Take a look at our complete list of pGreenFire 2.0 Lentivectors to see if we've already built the reporter you need.
pGreenFire 2.0 Lentivectors
| Catalog # | Lentivector | TRE |
|---|---|---|
| TR410PA/VA-P | pGreenFire 2.0-CMV positive control (pGF2-CMV-rFluc-T2A-GFP-mPGK-Puro) | Positive control |
| TR411PA/VA-P | pGreenFire 2.0-mCMV cloning & negative control (pGF2-mCMV-rFluc-T2A-GFP-mPGK-Puro) | Cloning & negative control |
| TR412PA/VA-P | pGreenFire 2.0 NFkB reporter (pGF2-NFκB-rFluc-T2A-GFP-mPGK-Puro) | NFκB |
| TR451PA/VA-P | pGreenFire 2.0 NFAT reporter (pGF2-NFAT-rFluc-T2A-GFP-mPGK-Puro) | NFAT |
| TR413PA/VA-P | pGreenFire 2.0 TCF/LEF reporter (pGF2-TCF/LEF-rFluc-T2A-GFP-mPGK-Puro) | TCF/LEF |
| TR452PA/VA-P | pGreenFire 2.0 AP-1 reporter (pGF2-AP1-rFluc-T2A-GFP-mPGK-Puro) | AP-1 |
| TR426PA/VA-P | pGreenFire 2.0 HIF-1 reporter (pGF2-HIF1-rFluc-T2A-GFP-mPGK-Puro) | HIF-1 |
| TR455PA/VA-P | pGreenFire 2.0 Estrogen response element reporter (pGF2-ERE-rFluc-T2A-GFP-mPGK-Puro) | Estrogen response element |