EXOSOME LIPIDOMICS & METABOLOMICS SERVICES
SBI’s Exosome Lipidomics & Metabolomics Services make it easy to expand your biomarker research to include exosomal lipids and other metabolites
Take your biomarker studies to the next level with exosomal lipidomics/metabolomics
What can lipidomics of exosomes tell you?
Lipids are an important part of cellular physiology, and are increasingly being recognized for their importance in exosome biology as well. Exosomes were recently shown to have the highest lipid-to-protein ratio of all classes of extracellular vesicles1, with lipid content that both differs from the parent cell the vesicles are shed from2 and also changes as exosomes undergo a variety of physiological processes3. These unique lipid profiles can serve as novel circulating biomarkers, and recent evidence suggests that specific lipid species carried by exosomes can also modulate the function of recipient cells4.
With so much information revealed by lipid content, lipidomics studies of exosomes are a great way to identify lipid-based biomarkers and for understanding vesicle biogenesis and function5.
What can metabolomics of exosomes tell you?
In addition to lipids, exosomes carry a wide range of metabolites that can also be used for biomarker discovery. A recent study found distinctly different populations of metabolites in PANC1 cells treated with TGF-β, a known driver of cancer progression, versus a control group, suggesting that metabolite profiling can be used for differentiating cellular states6. Like exosomal lipids, exosomal metabolites can also take an active role in the target cell, and have been shown to reprogram metabolic machinery upon uptake by cancer cells, fueling growth7.
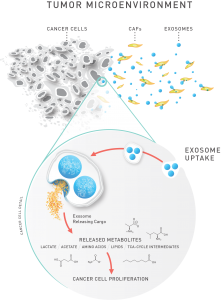
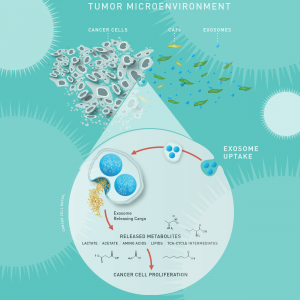
Figure 1. Exosomes affect metabolism in cancer cells. A recent study by Zhao, et al,7 demonstrated that exosomes from patient-derived cancer-associated fibroblasts (CAFs) can reprogram the cellular machinery in cancer cells. They used a metabolomics approach to demonstrate that exosomes shed from CAFs contain intact metabolites that fuel tumor growth. In addition, they showed that uptake of CAF exosomes inhibited mitochondrial oxidative phosphorylation, increasing glycolysis and glutamine-dependent reductive carboxylation.
Lipidomics & Metabolomics Service Highlights:
DISCOVER NOVEL CIRCULATING BIOMARKERS
LEARN MORE ABOUT EXOSOME BIOLOGY
SEND US YOUR SAMPLE AND RECEIVE DATA IN 4–6 WEEKS
Data deliverables:
- MS data will be delivered as Excel files with putative identification based on the m/z ratio of the analytes
- Differential analysis of lipids and metabolites between treatment groups will be included in the Excel files
How it Works
Our standard service offers UPLC-FTMS analysis of very polar metabolites in the aqueous phase, with lipids and other metabolites in the organic phase. Non-polar analysis also available upon request.
Note that all IDs are based only on retention time, accurate mass measurement, and preliminary MS/MS data, and cannot be guaranteed. Not all peaks will be identified, and we recommend that any peaks of interest—either due to the identity of the compound reported and/or change in intensity between samples—MUST be verified by additional targeted analysis at additional cost.
Input sample requirements
| BIOFLUID | VOLUME |
|---|---|
| Serum | 500μl–1ml |
| Plasma | 500μl–1ml |
| Cell Media | 5ml–10ml |
| Urine | 5ml–10ml |
| Spinal Fluid | 5ml–10ml |
| Ascites Fluid | 500μl–1ml |
| Other | Inquire |
Supporting Data
Successful genome engineering with SBI
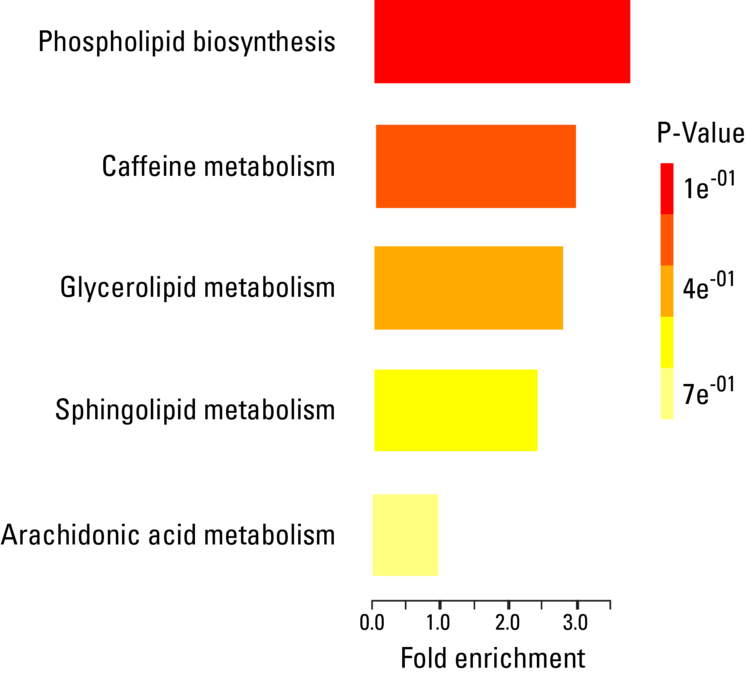
Figure 1. Successful genome engineering with SBI—effective, efficient knockout of miR-21 in HEK293 cells. gRNA, HR Donor design (with Puro and GFP selection markers), implementation, and analysis performed by SBI’s genome engineering services team. (A) Low relative levels of miR-21—as measured by qPCR in GFP-positive clones—demonstrate the effectiveness of the approach. (B) After excision with Cre recombinase, the inserted GFP and Puro markers are efficiently excised, leaving only a single LoxP site from the HR Donor. From Ho, TT, et al. Targeting noncoding RNAs with the CRISPR/Cas9 system in human cell lines. Nucleic Acids Res. 2015 Feb 18; 43(3):e17. PMCID: PMC4330338.
References
- Osteikoetxea X, et al. Improved characterization of EV preparations based on protein to lipid ratio and lipid properties. PLoS One. 2015;10(3):e0121184. PMCID: PMC4370721.
- Llorente A, et al. Molecular lipidomics of exosomes released by PC-3 prostate cancer cells. Biochim Biophys Acta. 2013;1831(7):1302-9. PMID: 24046871.
- Carayon K, et al. Proteolipidic composition of exosomes changes during reticulocyte maturation. J Biol Chem. 2011;286(39):34426-39. PMCID: PMC3190795.
- Subra C, et al. Exosomes account for vesicle-mediated transcellular transport of activatable phospholipases and prostaglandins. J Lipid Res. 2010;51(8):2105-20. PMCID: PMC2903822.
- Subra C, et al. Exosome lipidomics unravels lipid sorting at the level of multivesicular bodies. Biochimie. 2007;89(2):205-12. PMID: 17157973.
- Altadill T, et al. Enabling Metabolomics Based Biomarker Discovery Studies Using Molecular Phenotyping of Exosome-Like Vesicles. PLoS One. 2016 Mar 14;11(3):e0151339. PMCID: PMC4790956.
- Zhao H, et al. Tumor microenvironment derived exosomes pleiotropically modulate cancer cell metabolism. Elife. 2016 Feb 27;5. pii: e10250.&nb