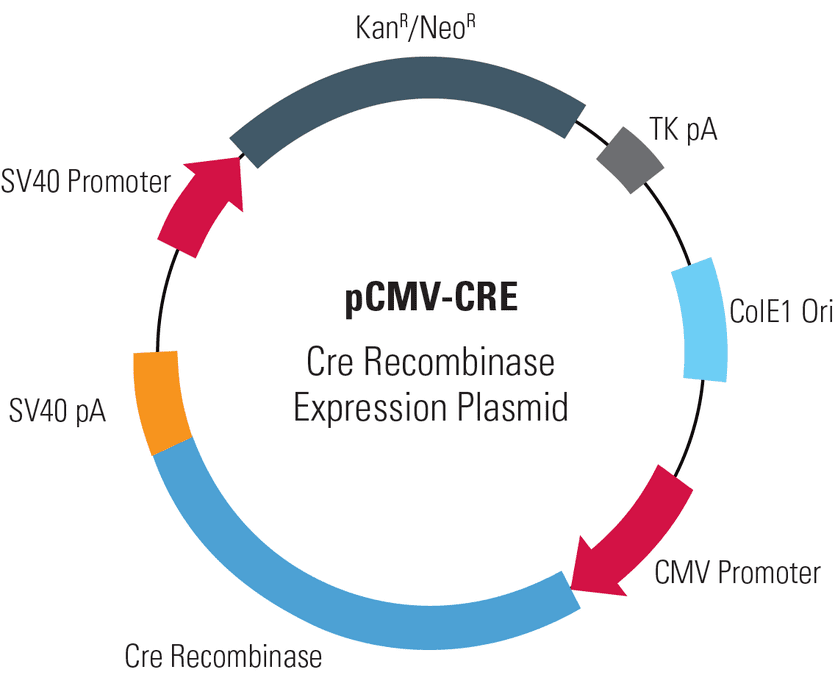
pCMV-CRE, Cre Recombinase Expression Plasmid
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| CRE100A-1 | Cre Recombinase Expression Plasmid (pCMV-CRE) | 10 µg | $683 |
|
||||
Overview
Overview
Get rid of selection cassettes and other unwanted sequences in your genome engineering projects
With the Cre-LoxP site-specific recombination system used in most of our HR Targeting Vectors, you can get rid of extra sequences, such as selection cassettes, by transiently expressing the Cre recombinase from the pCMV-CRE Cre Recombinase Expression Plasmid.
References
How It Works
How It Works
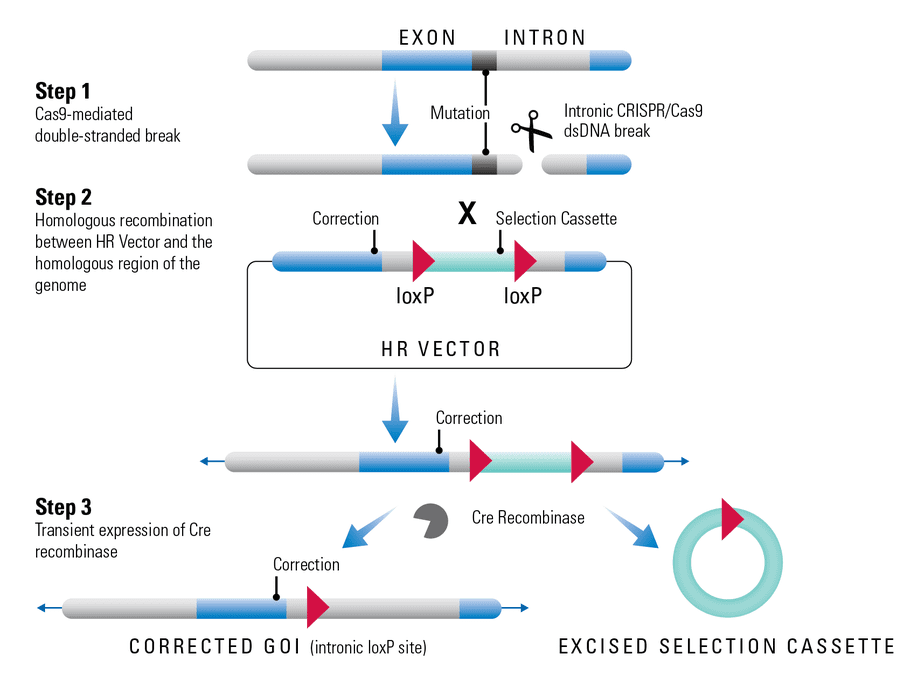
Removing a selection cassette with Cre recombinase
Cre recombinase catalyzes site-specific recombination between two LoxP sites. After recombination occurs, the sequences between the two LoxP sites are excised from the genome and a single LoxP site remains (Figure 1). Thus, with our HR Targeting Vectors that contain two LoxP sites, all the vector sequences that lie between the LoxP sites can be removed through transient expression of the Cre recombinase (Figure 1).
Figure 1. Removing a selection cassette using Cre-LoxP. The example shown demonstrates Cre-LoxP excision during genome editing but applies to any application where regions of an HR Targeting Vector with LoxP sites are integrated into the genome. In Step 1, Cas9 activity leads to a double-stranded break (DSB) in the genome. In Step 2, the regions of an HR Targeting Vector (also called an HR donor vector) that lie between the two homologous arms become integrated into the genome via homologous recombination. This integrated region typically contains a selection cassette with flanking LoxP sites to simplify finding clones where genome editing has occurred. In Step 3, the selection cassette is removed by transient expression of Cre recombinase.
Supporting Data
FAQs
Documentation
Citations
Related Products
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| CRE100A-1 | Cre Recombinase Expression Plasmid (pCMV-CRE) | 10 µg | $683 |
|
||||
Overview
Overview
Get rid of selection cassettes and other unwanted sequences in your genome engineering projects
With the Cre-LoxP site-specific recombination system used in most of our HR Targeting Vectors, you can get rid of extra sequences, such as selection cassettes, by transiently expressing the Cre recombinase from the pCMV-CRE Cre Recombinase Expression Plasmid.

References
How It Works
How It Works
Removing a selection cassette with Cre recombinase
Cre recombinase catalyzes site-specific recombination between two LoxP sites. After recombination occurs, the sequences between the two LoxP sites are excised from the genome and a single LoxP site remains (Figure 1). Thus, with our HR Targeting Vectors that contain two LoxP sites, all the vector sequences that lie between the LoxP sites can be removed through transient expression of the Cre recombinase (Figure 1).
Figure 1. Removing a selection cassette using Cre-LoxP. The example shown demonstrates Cre-LoxP excision during genome editing but applies to any application where regions of an HR Targeting Vector with LoxP sites are integrated into the genome. In Step 1, Cas9 activity leads to a double-stranded break (DSB) in the genome. In Step 2, the regions of an HR Targeting Vector (also called an HR donor vector) that lie between the two homologous arms become integrated into the genome via homologous recombination. This integrated region typically contains a selection cassette with flanking LoxP sites to simplify finding clones where genome editing has occurred. In Step 3, the selection cassette is removed by transient expression of Cre recombinase.